Description
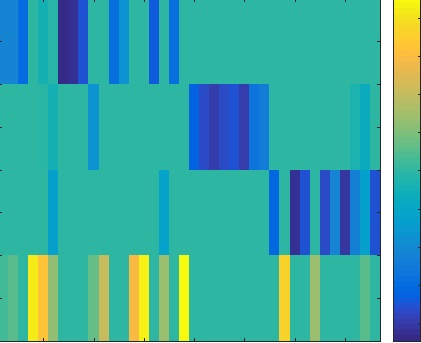
In this project, we proposed to use the penalized matrix decomposition (PMD) to extract metasamples from gene expression data. With the sparsity constrain on the decomposition factors, the extracted metasamples can well capture the intrinsic structures of the samples in the same class. Meanwhile, the PMD factors of each sample are good indicators of the class label of it. Compared with traditional methods, such as HC, SOM, AP, SC and NMF, the proposed method can identify the samples with complex classes. The experimental results on four representative data sets showed that the proposed method is able to effectively discover biological phenotypes, verifying that PMD is a powerful tool for gene expression data clustering.
It should be mentioned that we found experimentally that d can be used to determine the number of clusters according to its changing tendency. When d falls significantly from k to k+1, this means that all the meaningful patterns can be extracted using k clusters. If d has a gradual decay from k to k+1, this implies that more than k meaningful patterns may exist in the dataset. This is the reason that why d can be used to determine the number of clusters according to its changing tendency
However, at present, we can not conclude in theory that d must be or must not be the indicative value. It needs more investigation in the future. Fortunately, there have been some similar works on the statistical significance of matrix eigenvalues , which may be useful to our future study.

Relaxed averaged alternating reflection (RAAR) phase retrieval
http://pubs.broadinstitute.org/mpr/projects/NMF/NMF_final_supplement.pdf


Reviews
There are no reviews yet.