Matching of Micro-CT image With Histology
Image registration is an emerging field in medical research which is concerned with aligning volumetric data from multiple modalities, such as MRI, CT, ultrasound, and even histology. While registration algorithms for the alignment of volumes are well studied and documented, there has been little work on registration between 2D and 3D (Crum, Hartkens, & Hill, 2004). For the purpose of this study, a manual method was implemented to ‘search’ through the micro-CT volume for corresponding cross-section to histology.
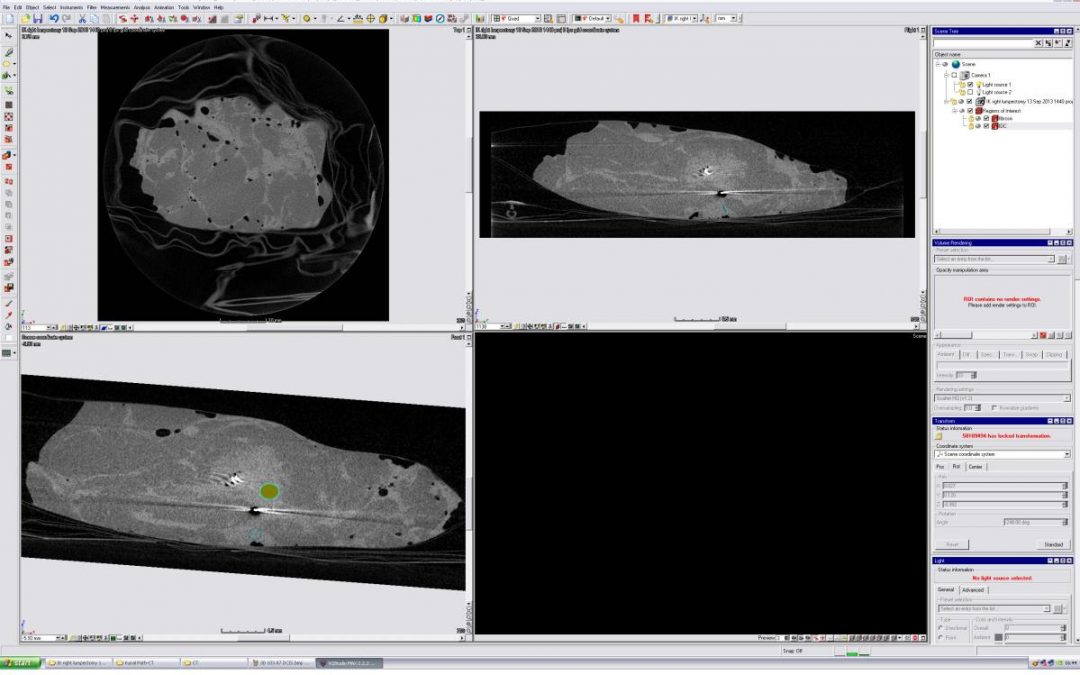
The micro-CT volume was loaded into the 3D analytical software package VGStudio MAX, a comprehensive software suite which allows for real-time coordinate transformations of micro-CT volumes, including translation and rotation, while providing views of CT orthoslices. On a separate screen, the pathology image to be matched was displayed. The micro-CT volume was rotated and manipulated, and corresponding orthoslices were scrolled through until the best matching slice was found.
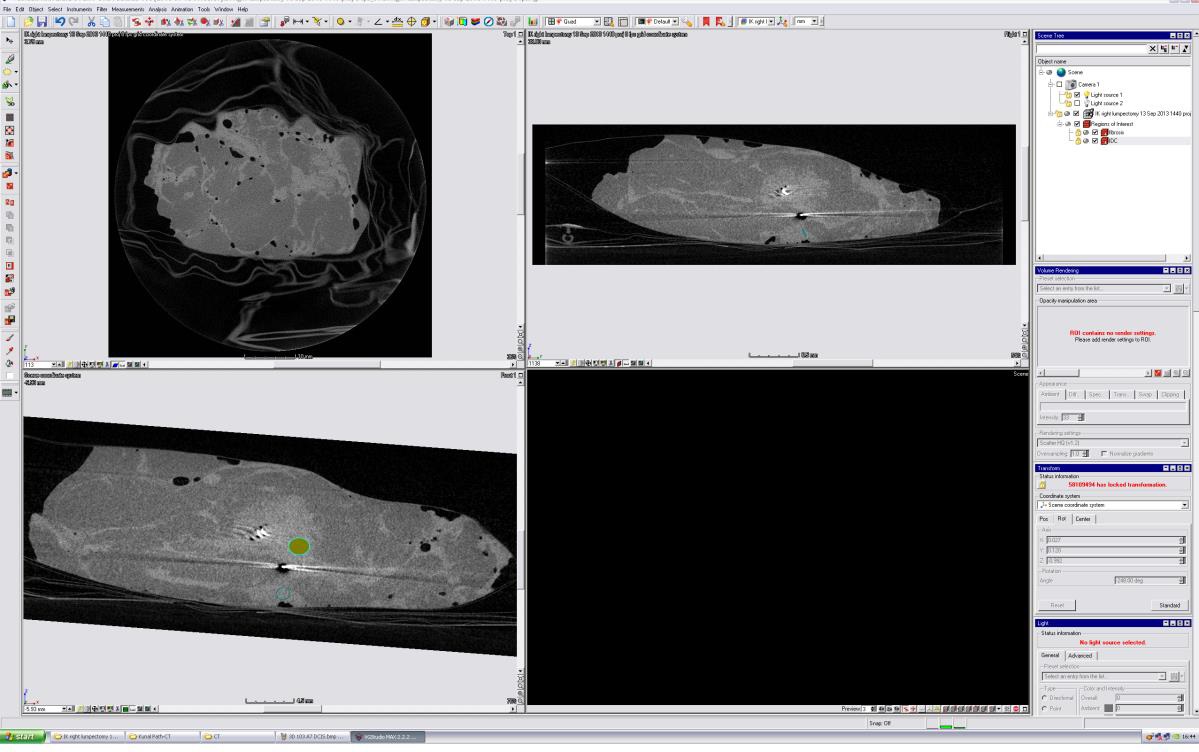
VGStudio Volumetric Analysis Suite
The determination of the “best “slice was a subjective one, but the location was confirmed by gross descriptions of submitted cassettes on the final pathology report, which cited landmarks such as the presence of a localization wire or a biopsy clip. Terms in the gross description such as “sequentially submitted” and “submitted from one end margin to the other” suggested that a series of slides would be matched along the same, or similar axis.